Syntax Export
Onyx supports exports of models to syntax specifications of other languages. This way, Onyx can be used for graphical model specification and other programs can then be used if their more advanced features are needed for estimating models (e.g., other estimators or non-normally distributed variables, etc.). To view model syntax, right-click a model and choose “Show Script” and select a syntax of your choice. Importantly, this includes the widely used R packages OpenMx and lavaan but also commercial software such as Mplus:
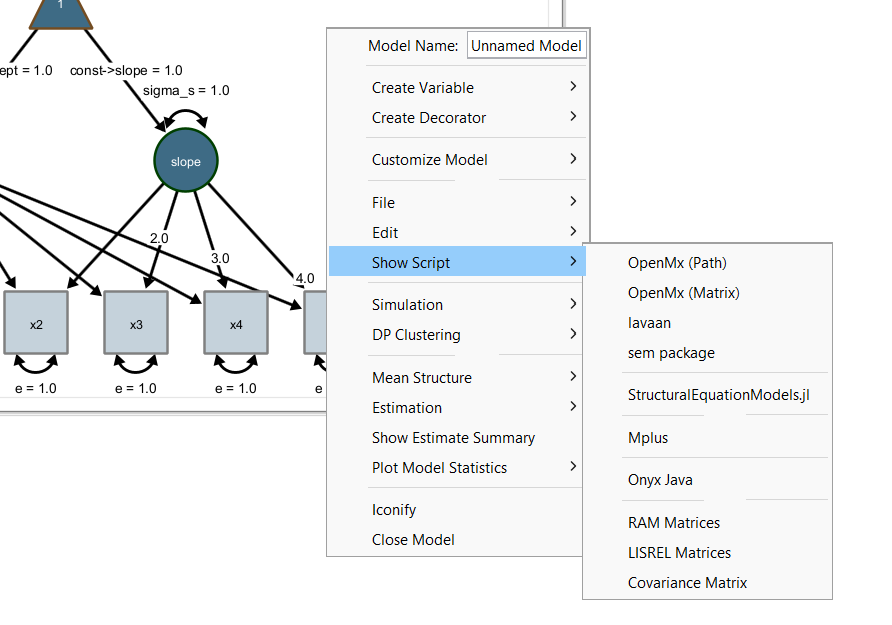
Importantly, the model syntax view remains linked to the model, that is, if the model graph is changed (e.g., by adding or removing variables or paths in the path diagram), the model syntax is updated. Onyx highlights newly added code parts in yellow.
Exercise
Create a linear latent growth curve model using the linear latent growth curve model wizard (right-click the desktop, select “Create new model -> Create new LGCM”; make sure to estimate a covariance between intercept and slope (click the respective option in the dialog)
Don’t forget to use some of the visual options to change the appearance of the diagram
Load the data “lgcm_simulated.csv” and obtain parameter estimates; inspect model fit
Open the model syntax view for lavaan
If you are familiar with lavaan and R: Copy the lavaan code to an R session; adapt the code to load the data file; estimate the model in lavaan; compare the results between Onyx and lavaan
Solution
The path diagram should look like this (showing the starting values; not the parameter estimates):
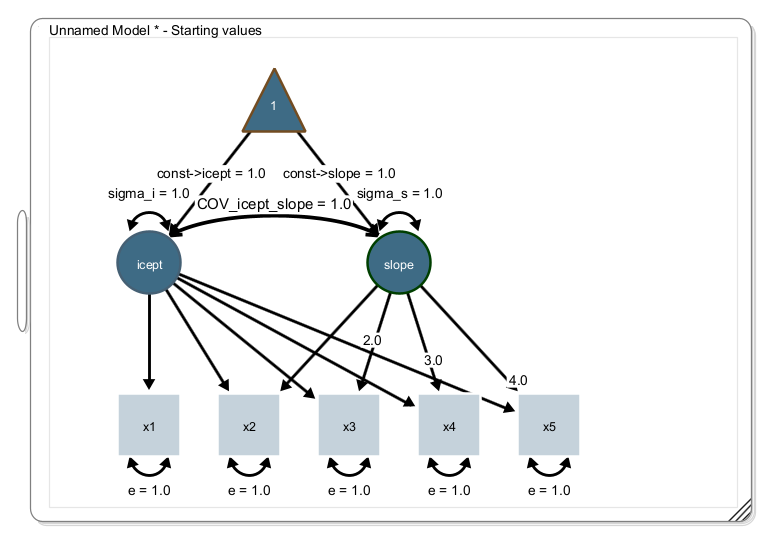
Load the dataset, select all variables, right-click on the data view and click “send data to model”. Model estimation starts and your parameter estimates and model fit statistics should look like this:
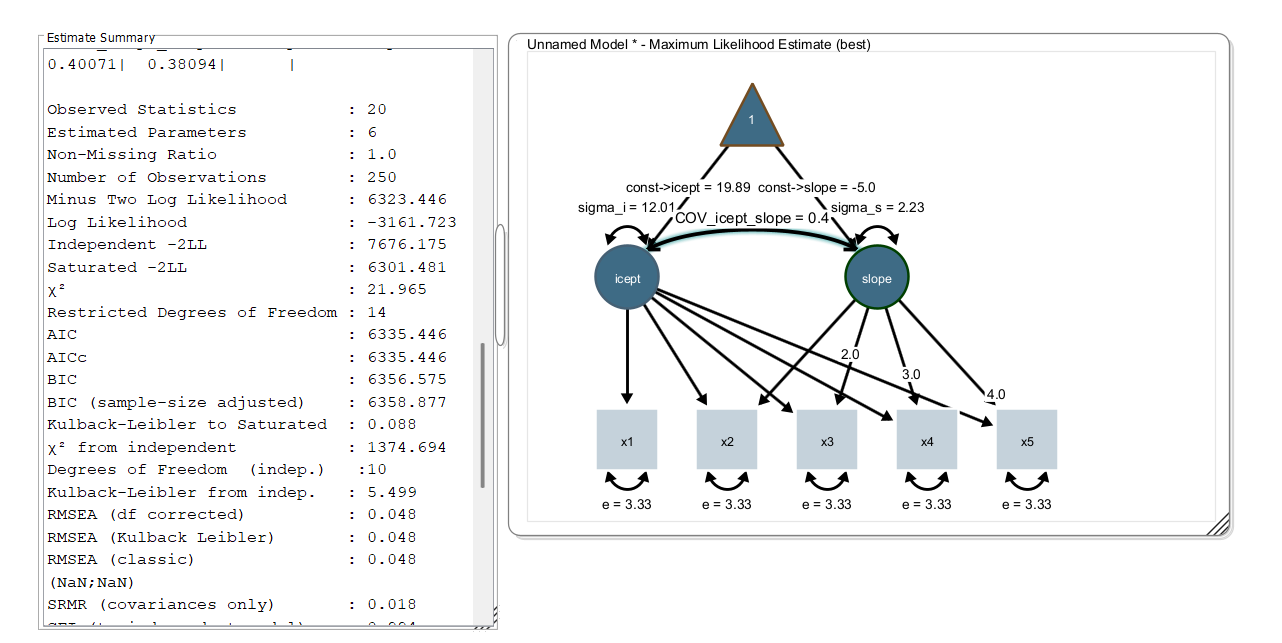 This is the lavaan model syntax as generated by Onyx:
This is the lavaan model syntax as generated by Onyx:
#
# This model specification was automatically generated by Onyx
#
library(lavaan);
modelData <- read.table(DATAFILENAME, header = TRUE) ;
model<-"
! regressions
icept=~1.0*x1
icept=~1.0*x2
slope=~1.0*x2
icept=~1.0*x3
slope=~2.0*x3
icept=~1.0*x4
slope=~3.0*x4
icept=~1.0*x5
slope=~4.0*x5
! residuals, variances and covariances
x1 ~~ e*x1
x2 ~~ e*x2
x3 ~~ e*x3
x4 ~~ e*x4
x5 ~~ e*x5
icept ~~ sigma_i*icept
slope ~~ sigma_s*slope
icept ~~ COV_icept_slope*slope
! means
icept ~ const__icept*1
slope ~ const__slope*1
x1 ~ 0.0*1
x2 ~ 0.0*1
x3 ~ 0.0*1
x4 ~ 0.0*1
x5 ~ 0.0*1
"
result<-lavaan(model, data=modelData, fixed.x=FALSE, missing="FIML")
summary(result, fit.measures=TRUE)
In this code, you only need to replace the placeholder “DATAFILENAME” with the path to a data file on your computer. Then, model estimation in R using the lavaan package yields:
Warning: Paket 'lavaan' wurde unter R Version 4.5.3 erstellt
This is lavaan 0.6-21
lavaan is FREE software! Please report any bugs.
lavaan 0.6-21 ended normally after 52 iterations
Estimator ML
Optimization method NLMINB
Number of model parameters 10
Number of equality constraints 4
Number of observations 250
Number of missing patterns 1
Model Test User Model:
Test statistic 21.965
Degrees of freedom 14
P-value (Chi-square) 0.079
Model Test Baseline Model:
Test statistic 1374.694
Degrees of freedom 10
P-value 0.000
User Model versus Baseline Model:
Comparative Fit Index (CFI) 0.994
Tucker-Lewis Index (TLI) 0.996
Robust Comparative Fit Index (CFI) 0.994
Robust Tucker-Lewis Index (TLI) 0.996
Loglikelihood and Information Criteria:
Loglikelihood user model (H0) -3161.723
Loglikelihood unrestricted model (H1) -3150.741
Akaike (AIC) 6335.446
Bayesian (BIC) 6356.575
Sample-size adjusted Bayesian (SABIC) 6337.554
Root Mean Square Error of Approximation:
RMSEA 0.048
90 Percent confidence interval - lower 0.000
90 Percent confidence interval - upper 0.084
P-value H_0: RMSEA <= 0.050 0.497
P-value H_0: RMSEA >= 0.080 0.076
Robust RMSEA 0.048
90 Percent confidence interval - lower 0.000
90 Percent confidence interval - upper 0.084
P-value H_0: Robust RMSEA <= 0.050 0.497
P-value H_0: Robust RMSEA >= 0.080 0.076
Standardized Root Mean Square Residual:
SRMR 0.014
Parameter Estimates:
Standard errors Standard
Information Observed
Observed information based on Hessian
Latent Variables:
Estimate Std.Err z-value P(>|z|)
icept =~
x1 1.000
x2 1.000
slope =~
x2 1.000
icept =~
x3 1.000
slope =~
x3 2.000
icept =~
x4 1.000
slope =~
x4 3.000
icept =~
x5 1.000
slope =~
x5 4.000
Covariances:
Estimate Std.Err z-value P(>|z|)
icept ~~
slope (COV_) 0.401 0.381 1.052 0.293
Intercepts:
Estimate Std.Err z-value P(>|z|)
icpt (cnst__c) 19.889 0.237 84.022 0.000
slop (cnst__s) -4.999 0.101 -49.371 0.000
.x1 0.000
.x2 0.000
.x3 0.000
.x4 0.000
.x5 0.000
Variances:
Estimate Std.Err z-value P(>|z|)
.x1 (e) 3.335 0.172 19.365 0.000
.x2 (e) 3.335 0.172 19.365 0.000
.x3 (e) 3.335 0.172 19.365 0.000
.x4 (e) 3.335 0.172 19.365 0.000
.x5 (e) 3.335 0.172 19.365 0.000
icept (sigm_) 12.007 1.257 9.551 0.000
slope (sgm_s) 2.230 0.230 9.699 0.000
As you can see, all parameter estimates and fit statistics are identical.


 This is the lavaan model syntax as generated by Onyx:
This is the lavaan model syntax as generated by Onyx: